Reproducibility of a Physiologically Based Pharmacokinetic and Pharmacodynamic (PBPK/PD) Model of Dapagliflozin
Abstract¶
A computational model in the form of a whole-body physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of dapagliflozin was developed to systematically evaluate the influence of patient-specific factors on drug disposition. Based on curated data from 28 clinical studies, the model simulates the absorption, distribution, metabolism and excretion (ADME) of the drug as well as its pharmacodynamics. The model accounts for variability in renal and hepatic function, and effects of food intake. The model is implemented in the Systems Biology Markup Language (SBML) standard. Analysis were performed utilizing the libroadrunner simulation library. Here, we demonstrate the computational reproducibility of the key findings from the primary publication, thereby verifying the consistency of the model implementation with the published results.
Introduction¶
In the primary publication Nemitz2025, a whole-body physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of dapagliflozin, an sodium-glucose co-transporter 2 (SGLT2) inhibitor prescribed for the management of type 2 diabetes mellitus (T2DM), was developed to mechanistically integrate factors driving variability. The model mechanistically integrates the key factors driving variability of dapagliflozin pharmacokinetics and pharmacodynamics, such as differences in renal Kasichayanula et al., 2011Kasichayanula et al., 2013 and/or hepatic Kasichayanula et al., 2011 function, and effects of food intake Kasichayanula et al., 2011. The model’s structure and parameters were derived from a comprehensive dataset curated from 28 published clinical studies. The dataset is available via the pharmacokinetics database PK-DB Grzegorzewski et al., 2021 and within the model files. The model’s development and scientific validation are described in detail in the primary paper Nemitz2025.
Here, we present the original model, encoded in the Systems Biology Markup Language (SBML) Hucka et al., 2019Keating et al., 2020, and the accompanying simulation scripts required to run the simulations and reproduce the key results presented in the primary publication.
Model Description¶
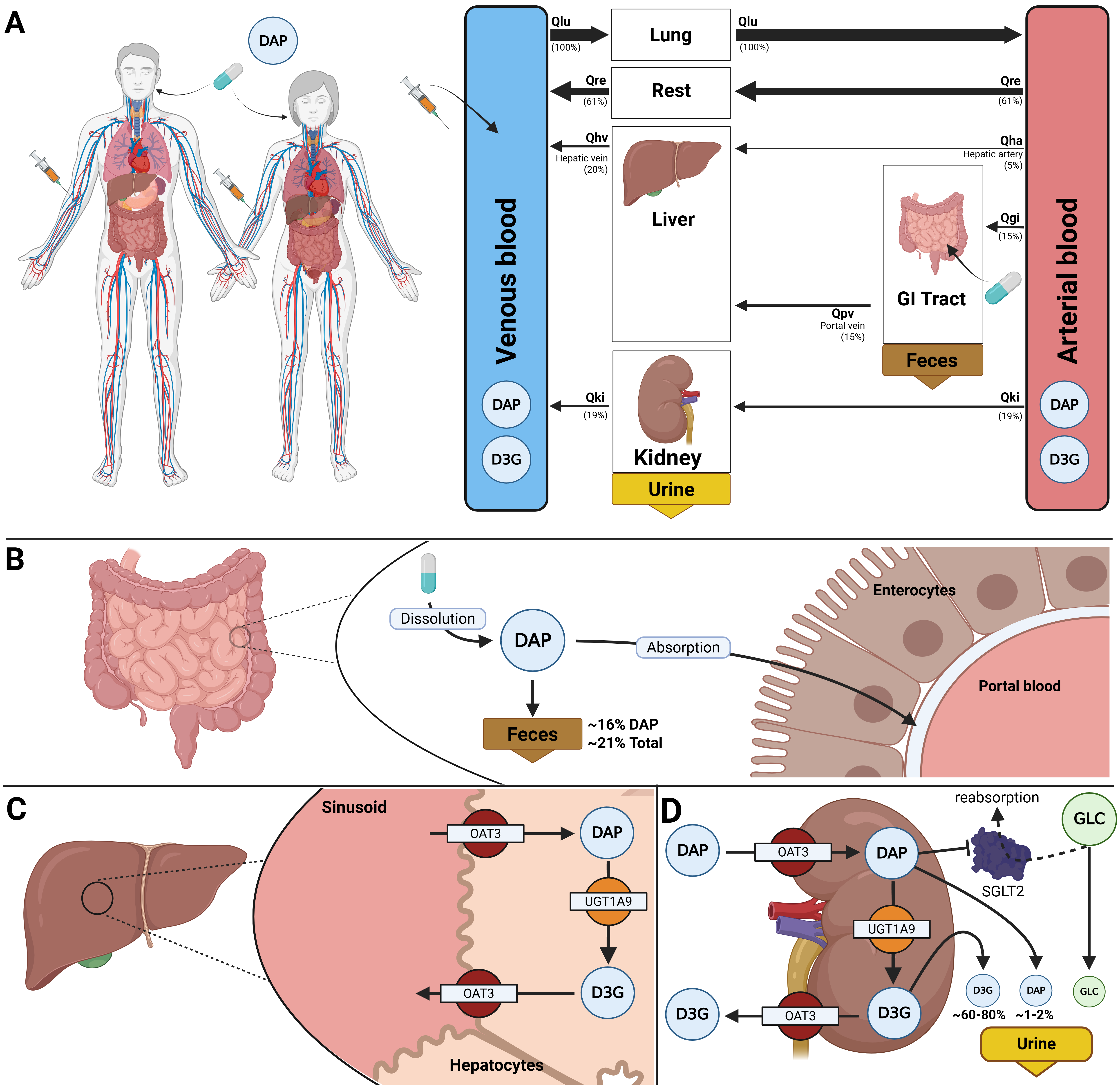
Figure 1:Whole-body PBPK/PD model of dapagliflozin and key factors influencing its disposition. A) Whole-body model illustrating dapagliflozin (DAP) administration (oral and intravenous), its systemic circulation via venous and arterial blood, and the key organs (liver, kidney, GI tract) involved in DAP metabolism, distribution, and excretion. B) Intestine model describing the absorption of DAP by enterocytes. DAP has a high bioavailability, with about 16% excreted in feces. C) Hepatic model depicting the uptake of DAP by hepatocytes and its conversion by UGT1A9 into its primary metabolite dapagliflozin-3-O-glucuronide (D3G). D) Renal model showing the uptake and excretion of DAP and D3G in urine and metabolic conversion of dapagliflozin to D3G. In the kidneys, dapagliflozin inhibits SGLT2, resulting in reduced glucose reabsorption and increased urinary glucose excretion.
The disposition of dapagliflozin is described using a whole-body physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model. The model consists of interconnected compartments that simulate key organs involved in the drug’s absorption, distribution, metabolism, and excretion (ADME) as well as in its pharmacodynamics. The mathematical framework of the model consists of ordinary differential equations (ODEs). A schematic overview of the model structure is provided in Figure 1.
The model integrates the three main organ submodels, which are interconnected via systemic circulation. The gastrointestinal tract model simulates the dissolution of orally administered dapagliflozin, its subsequent first-order absorption, and its fecal excretion. The liver model describes the primary metabolic pathway, where dapagliflozin is converted to its major metabolite dapagliflozin-3-O-glucuronide (D3G). The kidney model implements the renal excretion of dapagliflozin, D3G, and glucose, as well as the conversion of dapagliflozin to D3G. The pharmacodynamic component links dapagliflozin concentration in the kidneys to urinary glucose excretion (UGE) through inhibition of renal glucose reabsorption. This process is affected by fasting plasma glucose and the renal threshold for glucose.
The model accounts for patient-specific factors through the corresponding scaling parameters. Renal impairment was modeled as a progressive decline in renal function by scaling the factor frenal. Hepatic impairment was modeled as a gradual increase in cirrhosis by scaling liver function with the parameter fcirrhosis. Food effect was incorporated via the intestinal absorption scaling factor fabsorption.
The PBPK/PD model and its tissue-specific submodels were developed using the Systems Biology Markup Language (SBML) Hucka et al., 2019Keating et al., 2020. Programming and visualization of the models were performed using the sbmlutils König, 2024 and cy3sbml König et al., 2012 libraries. Numerical solutions for the ordinary differential equations (ODEs) underlying the model were computed using sbmlsim König, 2021, which is powered by the high-performance SBML simulation engine libroadrunner Welsh et al., 2023Somogyi et al., 2015. The tissue submodels were developed as SBML submodels and coupled with the whole-body model using the hierarchical model composition (comp) SBML extension Smith et al., 2015. The complete model, submodels, reference simulations, and visualizations are available as a COMBINE archive (OMEX) Bergmann et al., 2014Bergmann et al., 2015. The model is annotated with extensive metadata using the open modeling and exchange (OMEX) metadata specification Neal et al., 2020Neal et al., 2019. The model was validated using the SBML validator, with the model passing all validation tests without errors or warnings. The FAIRness of the model was increased by following the FAIRification of computational models in the biological workflow Balaur et al., 2025.
The model and all associated materials (mathematical formulation, simulation scripts, parameters, and documentation) are publicly available in SBML format and OMEX archive under a CC-BY 4.0 license at https://
Computational Simulation¶
All simulations were performed using Python 3.13 together with the high-performance libRoadRunner simulation engine. The workflow was tested across multiple platforms, including Ubuntu 24.04/25.10 and Windows 11. For SBML model handling and simulation, we relied on the sbmlutils and sbmlsim libraries, while data management and figure generation were carried out with standard scientific Python packages.
To ensure reproducibility, we provide two equivalent setups for regenerating all figures presented in Section ??: (1) a local Python installation using uv, and (2) a containerized workflow using Docker. Both approaches reproduce all results from the primary publication. Reproducibility is continuously validated through automated integration tests, with results available at https://
Python with uv (local install)¶
This workflow installs the package directly on your machine using uv.
Prerequisite: uv must be installed on your system (https://
Clone the repository and move into its folder:
git clone https://github.com/matthiaskoenig/dapagliflozin-model.git
cd dapagliflozin-modelSet up the uv virtual environment and install all dependencies:
uv syncRun the complete analysis:
uv run run_dapagliflozin -a all -r resultsAll reproduced figures and outputs are written to ./results/ inside the repository.
Alternatively, you can use any other way to setup a local python environment (e.g. conda) and install the package after cloning the repository via:
pip install -e . or directly from the main branch via:
pip install git+https://github.com/matthiaskoenig/dapagliflozin-model.git@mainThe full analysis can be run in the python environment via:
(env) run_dapagliflozin -a all -r resultsDocker (containerized)¶
This workflow runs the analysis in a preconfigured Docker container.
Prerequisite: Docker must be installed on your system (https://
Start the container and mount a local results/ directory:
docker run -v "${PWD}/results:/results" -it matthiaskoenig/dapagliflozin:latest /bin/bashInside the container, run the analysis. Results will be written to the mounted folder:
uv run run_dapagliflozin -a all -r /resultsThe reproduced figures and outputs are then accessible on the host system in ./results/.
If file access is restricted on Linux due to permissions, adjust ownership and rights as follows:
sudo chown $(id -u):$(id -g) -R "${PWD}/results"
sudo chmod 775 "${PWD}/results"Available Options¶
Specific parts of the analysis can be executed by providing command-line arguments. A full overview of the available options is obtained via:
uv run run_dapagliflozin --helpOutputs¶
The workflow reproduces all figures and results from the primary publication, including:
All results are stored in the results/ directory. This directory contains the individual figure panels in PNG format as well as an automatically generated HTML report (index.html) that consolidates all figures into a single document. The content of this report directly corresponds to Figures 2–6 in the manuscript.
Reproducibility Goals¶
The reproducibility of the dapagliflozin PBPK/PD model was confirmed by reproducing key figures from the original publication and its supplementary material. The figures presented here are a selection chosen to demonstrate consistent reproduction of results across different dose levels, pathophysiological, and prandial states. Tables 1–2 provide an overview of the simulation observables and the parameter changes specific to each study, experiment, or scan. The model and simulation scripts can be used to reproduce the full set of results from the original study and its supplements.
Table 1:Plotted observables and parameter changes per study simulation (continued). Square brackets around SBML species ids indicate concentrations (amount/volume units). Square brackets enclosing numerical values indicate parameter ranges, whereas curly brackets indicate sets of discrete choices.
| StudyID | Plotted | Changes |
Boulton2013 | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Cho2021 | [Cve_dap] | [tl]BW = 72.8 kg |
[0.7em] FDAMB102002 FDA, 2013 | [Cve_dap], KI__UGE | [tl]PODOSE_dap mg |
[0.7em] FDAMB102003 FDA, 2013 | [Cve_dap], Aurine_dap, KI__UGE | [tl]PODOSE_dap mg |
[0.7em] FDAMB102006 FDA, 2013 | Aurine_daptot, Aurine_dap, Aurine_d3g, Afeces_daptot, Afeces_dap | [tl]PODOSE_dap = 50 mg |
[0.7em] FDAMB102007 FDA, 2013 | Aurine_dap, Aurine_d3g, KI__UGE | [tl]PODOSE_dap mg |
[0.7em] Gould2013 | KI__UGE | [tl]PODOSE_dap |
[0.7em] Hwang2022a | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Imamura2013 | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Jang2020 | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Kasichayanula2011 | [Cve_dap], [Cve_d3g] | [tl]PODOSE_dap = 10 mg |
[0.7em] Kasichayanula2011a | [Cve_dap], [Cve_d3g], KI__UGE | [tl]PODOSE_dap mg |
[0.7em] Kasichayanula2011b | [Cve_dap], [Cve_d3g] | [tl]PODOSE_dap = 10 mg |
[0.7em] Kasichayanula2011c | [Cve_dap] | [tl]PODOSE_dap mg |
[0.7em] Kasichayanula2012 | [Cve_dap] | [tl]PODOSE_dap = 20 mg |
[0.7em] Kasichayanula2013 | [Cve_dap], [Cve_d3g], Aurine_dap, Aurine_d3g | [tl]PODOSE_dap mg |
[0.7em] Kasichayanula2013a | [Cve_dap], [Cve_d3g], Aurine_dap, Aurine_d3g | [tl]PODOSE_dap = 10 mg |
[0.7em] Khomitskaya2018 | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Kim2023 | [Cve_dap], KI__UGE | [tl]PODOSE_dap = 10 mg |
[0.7em] Kim2023a | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Komoroski2009 | [Cve_dap], Aurine_dap, KI__UGE | [tl]PODOSE_dap |
[0.7em] LaCreta2016 | [Cve_dap] | [tl]PODOSE_dap mg |
[0.7em] Obermeier2010 | [Cve_dap], [Cve_daptot] | [tl]PODOSE_dap = 50 mg |
[0.7em] Sha2015 | [Cve_dap], KI__UGE, KI__RTG | [tl]PODOSE_dap = 10 mg |
[0.7em] Shah2019a | [Cve_dap] | [tl]PODOSE_dap = 5 mg |
[0.7em] vanderAartvanderBeek2020 | [Cve_dap] | [tl]PODOSE_dap = 10 mg |
[0.7em] Watada2019 | [Cve_dap], [Cve_d3g], KI__UGE | [tl]PODOSE_dap mg |
[0.7em] Yang2013 | [Cve_dap], [Cve_d3g], KI__UGE | [tl]PODOSE_dap mg |
Table 2:Plotted observables and parameter changes per simulation experiment (continued). Square brackets around SBML species ids indicate concentrations (amount/volume units). Square brackets enclosing numerical values indicate parameter ranges, whereas curly brackets indicate sets of discrete choices.
| Simulation | Plotted | Changes |
| DoseDependencyExperiment | [Cve_dap], [Cve_d3g], Aurine_dap, Aurine_d3g, Afeces_dap, KI__RTG, KI__UGE | [tl]PODOSE_dap mg |
[0.7em] FoodEffect | [Cve_dap], [Cve_d3g], Aurine_dap, Aurine_d3g, Afeces_dap, KI__RTG, KI__UGE | [tl]PODOSE_dap = 10 mg |
[0.7em] HepaticRenalImpairment | [Cve_dap], [Cve_d3g], Aurine_dap, Aurine_d3g, Afeces_dap, KI__RTG, KI__UGE | [tl]PODOSE_dap = 10 mg |
[0.7em] DapagliflozinParameterScan | [Cve_dap], [Cve_d3g], Aurine_dap, Aurine_d3g, Afeces_dap, KI__UGE, AUCinf, Cmax, thalf, UGE(24 hr) | [tl]f_cirrhosis (PODOSE_dap = 10 mg) |
Reproduction of Study Simulations¶
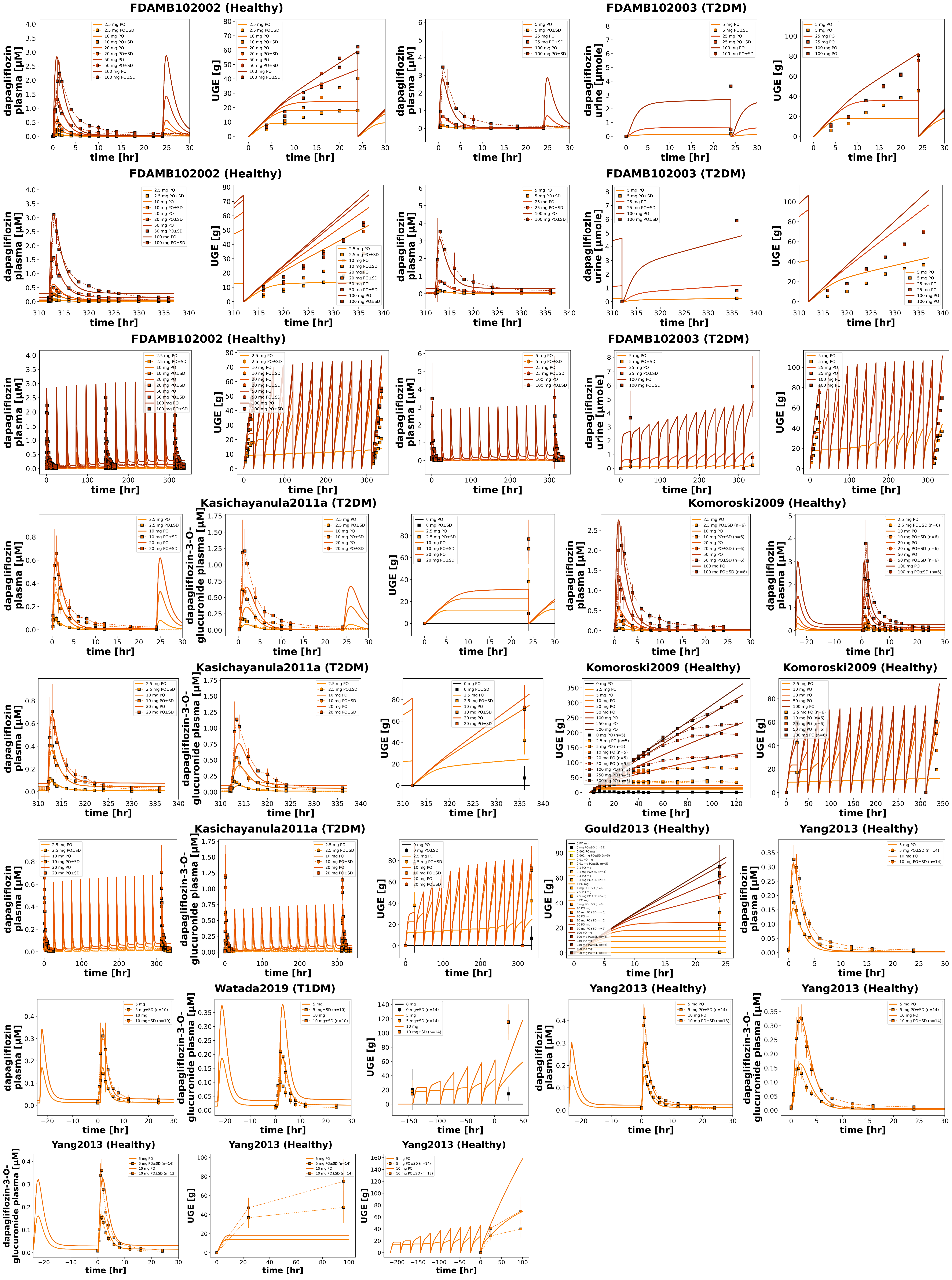
Figure 2:Reproduction of study simulations (dose dependency) from the primary publication. Data is taken from FDA (2013)FDA (2013)Gould et al. (2013)Kasichayanula et al. (2011)Komoroski et al. (2009)Watada et al. (2019)Yang et al. (2013).
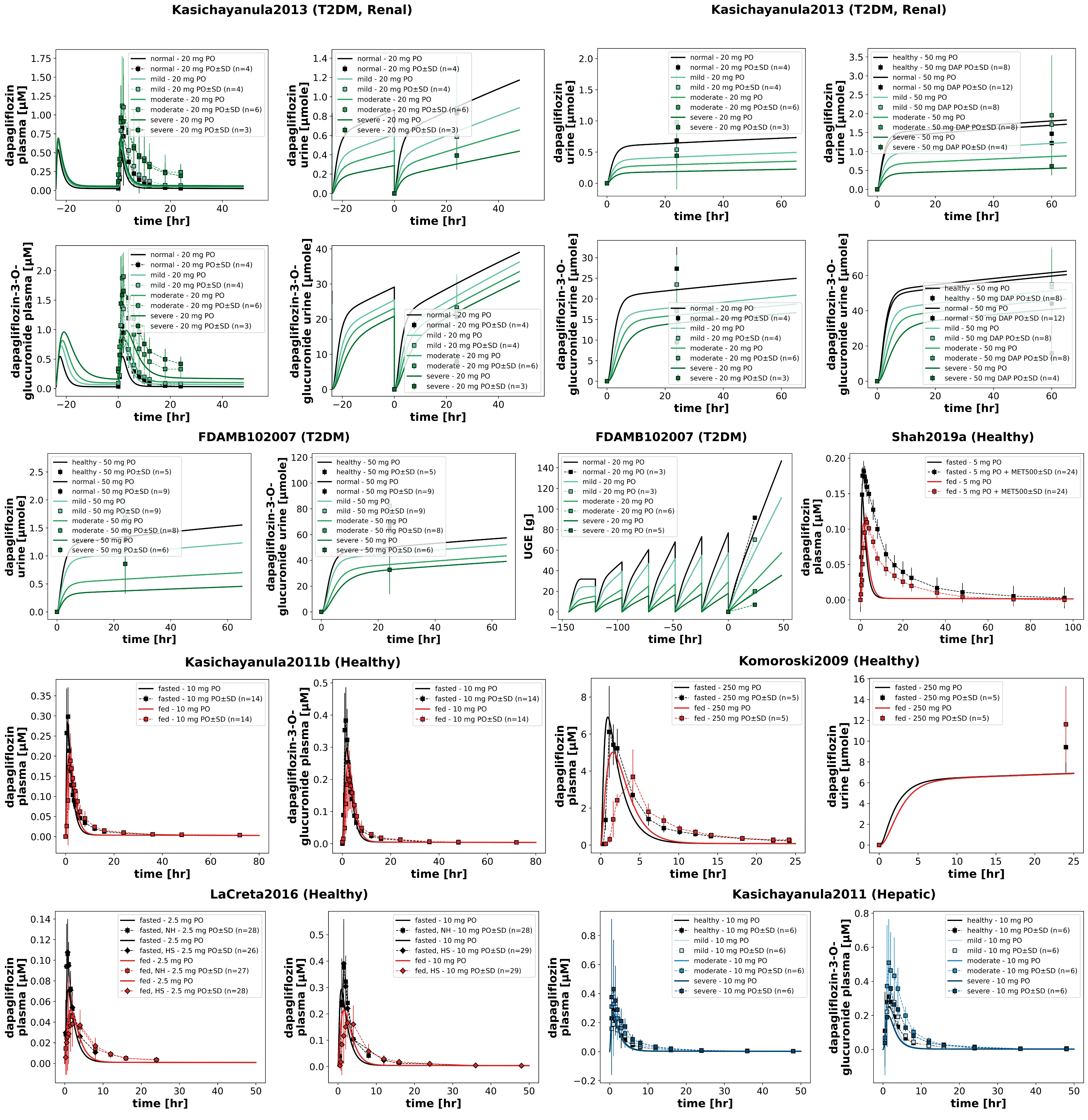
Figure 3:Reproduction of study simulations (renal impairment, hepatic impairment, and food effects) from the primary publication. Data is taken from FDA (2013)Kasichayanula et al. (2011)Kasichayanula et al. (2011)Kasichayanula et al. (2013)Komoroski et al. (2009)LaCreta et al. (2016)Shah et al. (2019).
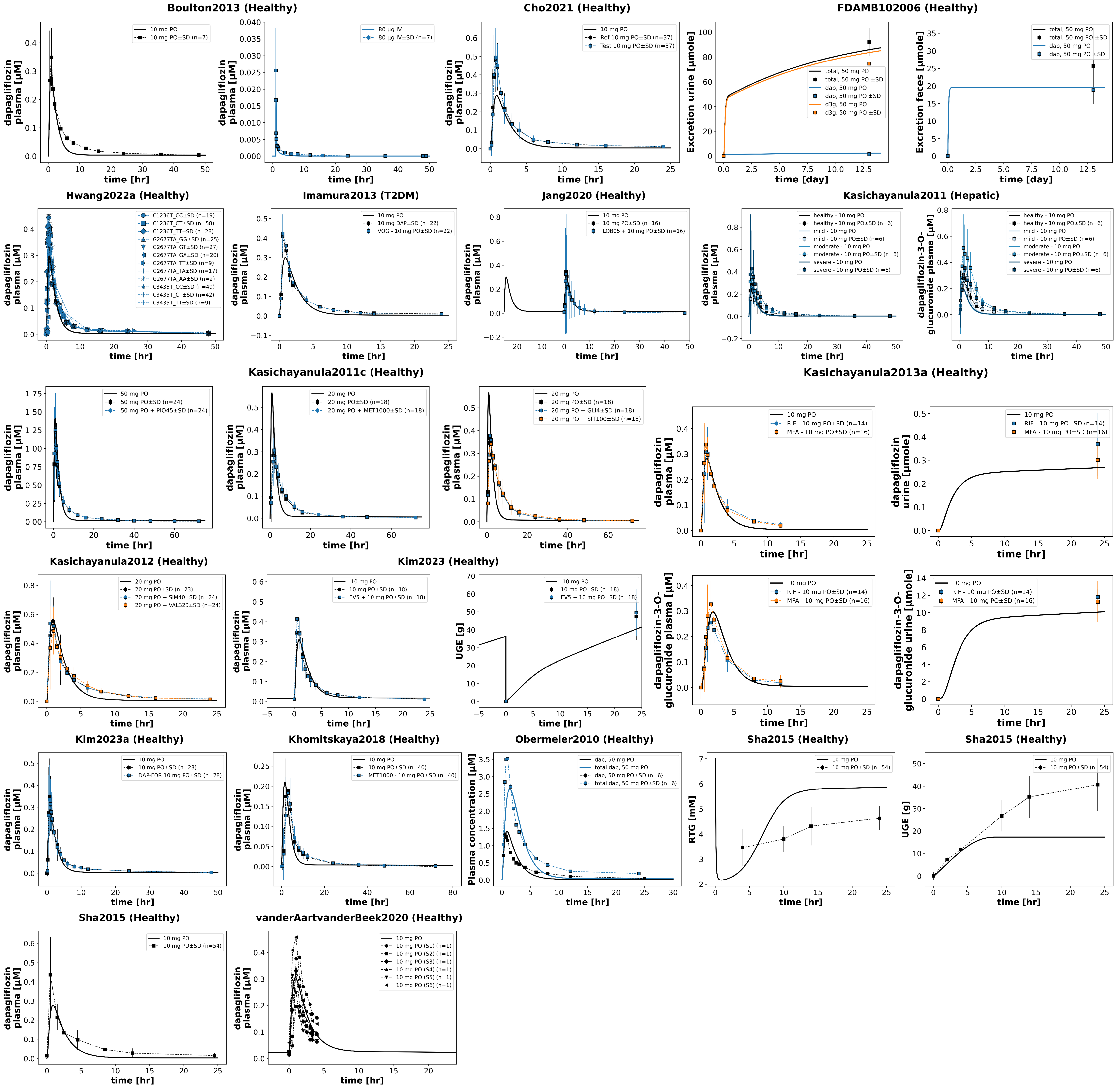
Figure 4:Reproduction of supplementary study simulations from the primary publication. Data is taken from Boulton et al. (2013)Cho et al. (2021)FDA (2013)Hwang et al. (2022)Imamura et al. (2013)Jang et al. (2020)Kasichayanula et al. (2011)Kasichayanula et al. (2011)Kasichayanula et al. (2012)Kasichayanula et al. (2013)Khomitskaya et al. (2018)Kim et al. (2023)Kim et al. (2023)Obermeier et al. (2010)Sha et al. (2015)van der Aart-van der Beek et al. (2020).
Reproduction of Simulations Experiments and Scans¶
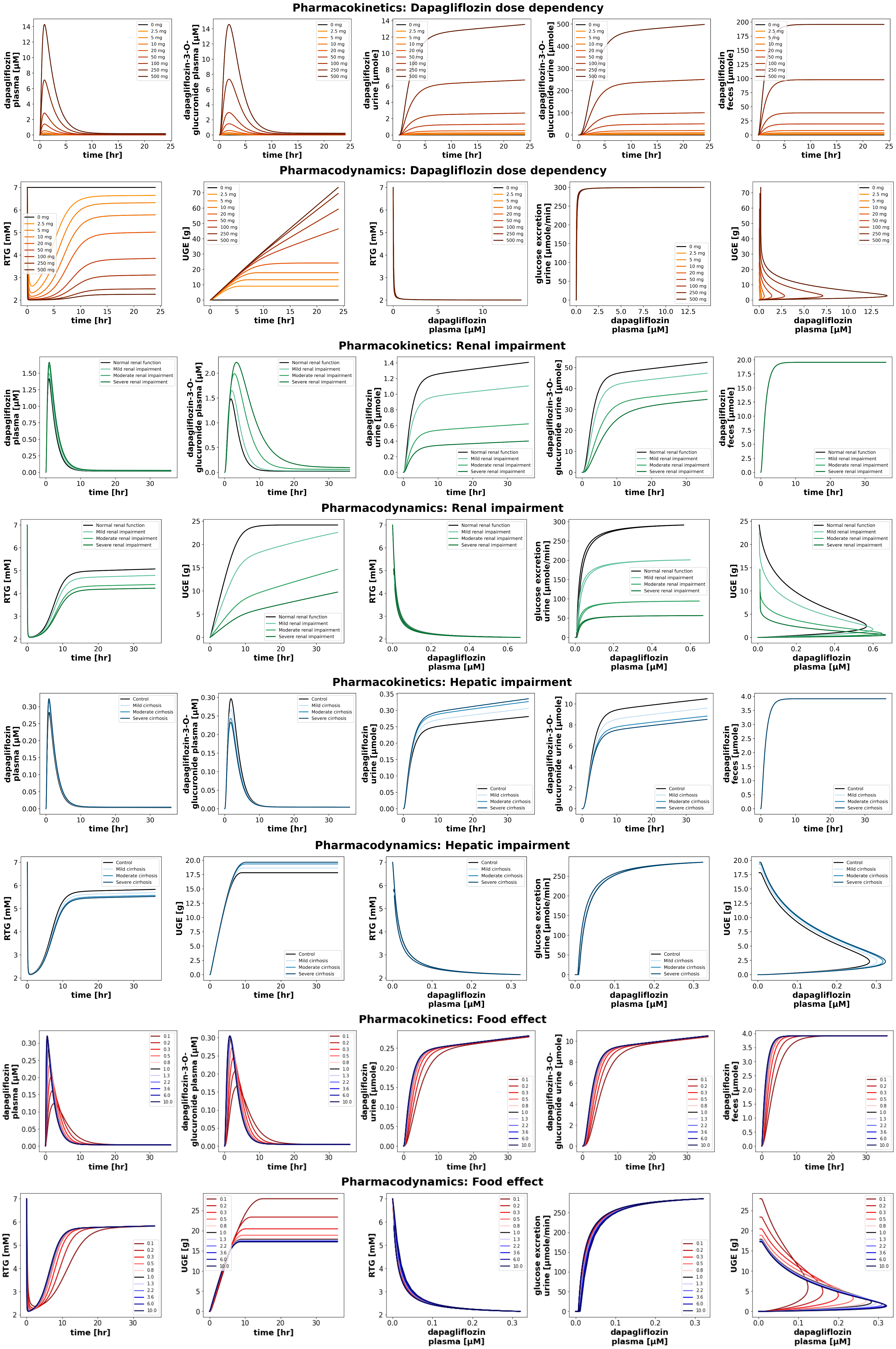
Figure 5:Reproduction of simulation experiments from the primary publication.
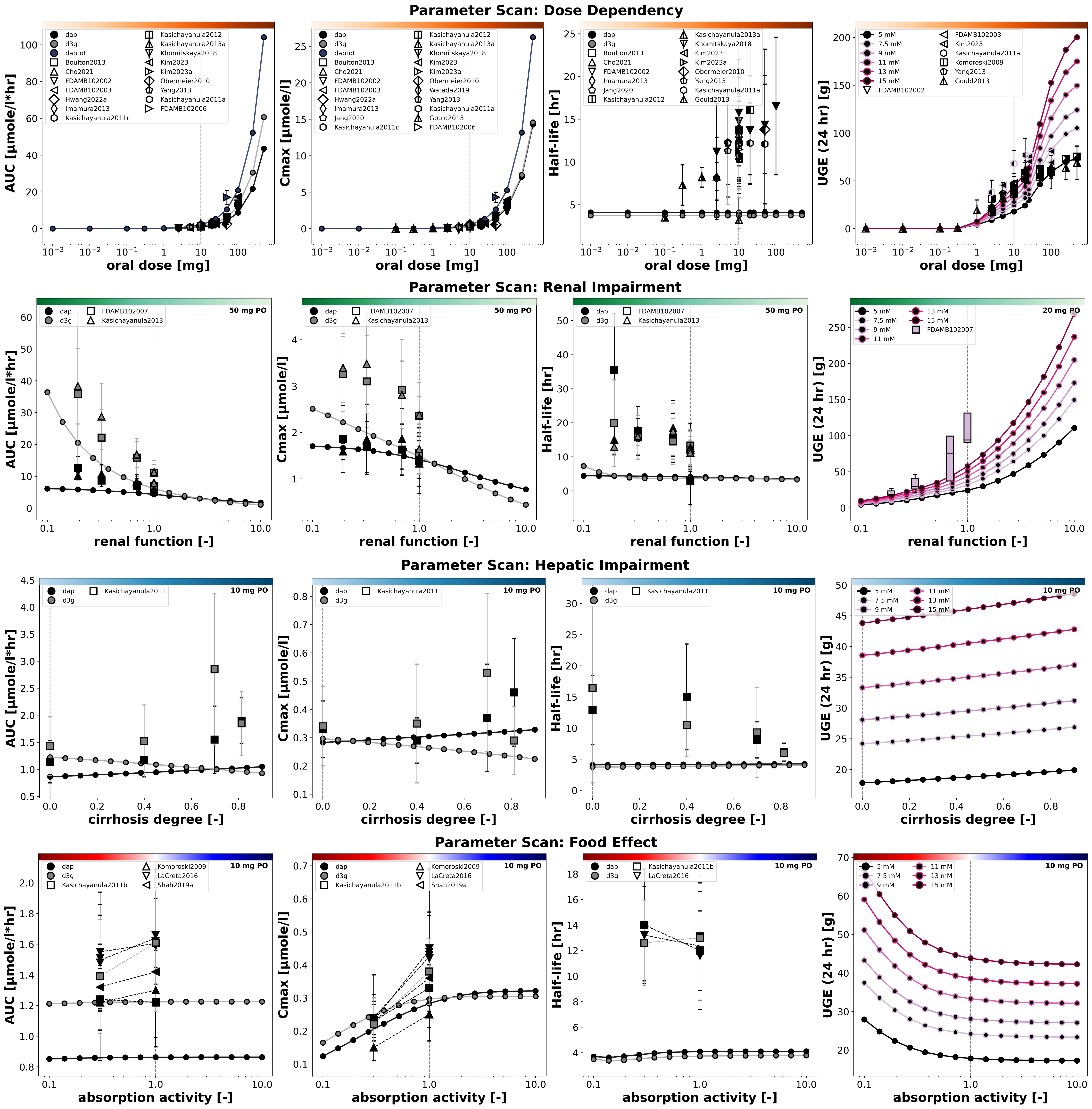
Figure 6:Reproduction of parameter scans from the primary publication. Data is taken from Boulton et al. (2013)Cho et al. (2021)FDA (2013)FDA (2013)FDA (2013)FDA (2013)Gould et al. (2013)Hwang et al. (2022)Imamura et al. (2013)Jang et al. (2020)Kasichayanula et al. (2011)Kasichayanula et al. (2011)Kasichayanula et al. (2011)Kasichayanula et al. (2011)Kasichayanula et al. (2012)Kasichayanula et al. (2013)Kasichayanula et al. (2013)Khomitskaya et al. (2018)Kim et al. (2023)Kim et al. (2023)Komoroski et al. (2009)LaCreta et al. (2016)Obermeier et al. (2010)Shah et al. (2019)Yang et al. (2013).
Discussion¶
We have demonstrated the computational reproducibility of the key findings from the dapagliflozin PBPK/PD model presented in the primary publication. Using the provided simulation scripts, all figures were regenerated without modifying parameters or structure, verifying the consistency of the model. Reproducibility was confirmed across different operating systems using both a local installation with uv and a Dockerized workflow. The uv-based approach allows users to install the package and dependencies natively, while the containerized workflow provides a fully preconfigured environment and ensures consistent results independent of the local setup. Encoding the model in SBML with hierarchical composition removes ambiguity and allows modular reuse of the tissue submodels. Together with the use of community standards and FAIR practices, this provides a transparent and reusable resource that can be applied or extended in future pharmacokinetic/pharmacodynamic modeling work.
Author Contributions¶
M.E., N.N. and M.K. contributed to conceptualization, methodology, data curation, and development of the PBPK/PD model. M.E., N.N., M.M., and M.K. contributed to analyses, software, and visualization. M.E. M.M., and M.K. contributed to the reproducibility of the computational workflow. M.E. wrote the original draft. M.E., N.N., M.M., and M.K. contributed to manuscript review and editing. M.K. provided supervision throughout the project. All authors approved the final manuscript.
Funding¶
Matthias König (MK) was supported by the Federal Ministry of Research, Technology and Space (BMFTR, Germany) within ATLAS by grant number 031L0304B and by the German Research Foundation (DFG) within the Research Unit Program FOR 5151 "QuaLiPerF (Quantifying Liver Perfusion-Function Relationship in Complex Resection - A Systems Medicine Approach)" by grant number 436883643 and by grant number 465194077 (Priority Programme SPP 2311, Subproject SimLivA). Mariia Myshkina was supported by the Federal Ministry of Research, Technology and Space (BMFTR, Germany) within ATLAS by grant number 031L0304B and by the German Research Foundation (DFG) within the Priority Programme SPP 2311, Subproject SimLivA by grant number 465194077. This work was supported by the BMFTR-funded de.NBI Cloud within the German Network for Bioinformatics Infrastructure (de.NBI) (031A537B, 031A533A, 031A538A, 031A533B, 031A535A, 031A537C, 031A534A, 031A532B).
Acknowledgments¶
Figures were created in BioRender. König, M. (2025) https://
- Kasichayanula, S., Chang, M., Hasegawa, M., Liu, X., Yamahira, N., LaCreta, F. P., Imai, Y., & Boulton, D. W. (2011). Pharmacokinetics and Pharmacodynamics of Dapagliflozin, a Novel Selective Inhibitor of Sodium-Glucose Co-Transporter Type 2, in Japanese Subjects without and with Type 2 Diabetes Mellitus. Diabetes, Obesity & Metabolism, 13(4), 357–365. 10.1111/j.1463-1326.2011.01359.x
- Kasichayanula, S., Liu, X., Pe Benito, M., Yao, M., Pfister, M., LaCreta, F. P., Humphreys, W. G., & Boulton, D. W. (2013). The Influence of Kidney Function on Dapagliflozin Exposure, Metabolism and Pharmacodynamics in Healthy Subjects and in Patients with Type 2 Diabetes Mellitus. British Journal of Clinical Pharmacology, 76(3), 432–444. 10.1111/bcp.12056
- Kasichayanula, S., Liu, X., Zhang, W., Pfister, M., LaCreta, F. P., & Boulton, D. W. (2011). Influence of Hepatic Impairment on the Pharmacokinetics and Safety Profile of Dapagliflozin: An Open-Label, Parallel-Group, Single-Dose Study. Clinical Therapeutics, 33(11), 1798–1808. 10.1016/j.clinthera.2011.09.011
- Kasichayanula, S., Liu, X., Zhang, W., Pfister, M., Reele, S. B., Aubry, A.-F., LaCreta, F. P., & Boulton, D. W. (2011). Effect of a High-Fat Meal on the Pharmacokinetics of Dapagliflozin, a Selective SGLT2 Inhibitor, in Healthy Subjects. Diabetes, Obesity & Metabolism, 13(8), 770–773. 10.1111/j.1463-1326.2011.01397.x
- Grzegorzewski, J., Brandhorst, J., Green, K., Eleftheriadou, D., Duport, Y., Barthorscht, F., Köller, A., Ke, D. Y. J., De Angelis, S., & König, M. (2021). PK-DB: Pharmacokinetics Database for Individualized and Stratified Computational Modeling. Nucleic Acids Research, 49(D1), D1358–D1364. 10.1093/nar/gkaa990
- Hucka, M., Bergmann, F. T., Chaouiya, C., Dräger, A., Hoops, S., Keating, S. M., König, M., Novère, N. L., Myers, C. J., Olivier, B. G., Sahle, S., Schaff, J. C., Sheriff, R., Smith, L. P., Waltemath, D., Wilkinson, D. J., & Zhang, F. (2019). The Systems Biology Markup Language (SBML): Language Specification for Level 3 Version 2 Core Release 2. Journal of Integrative Bioinformatics, 16(2). 10.1515/jib-2019-0021
- Keating, S. M., Waltemath, D., König, M., Zhang, F., Dräger, A., Chaouiya, C., Bergmann, F. T., Finney, A., Gillespie, C. S., Helikar, T., Hoops, S., Malik-Sheriff, R. S., Moodie, S. L., Moraru, I. I., Myers, C. J., Naldi, A., Olivier, B. G., Sahle, S., Schaff, J. C., … SBML Level 3 Community members. (2020). SBML Level 3: An Extensible Format for the Exchange and Reuse of Biological Models. Molecular Systems Biology, 16(8), e9110. 10.15252/msb.20199110
- König, M. (2024). Sbmlutils: Python Utilities for SBML. Zenodo. 10.5281/ZENODO.13325770
- König, M., Dräger, A., & Holzhütter, H.-G. (2012). CySBML: A Cytoscape Plugin for SBML. Bioinformatics, 28(18), 2402–2403. 10.1093/bioinformatics/bts432
- König, M. (2021). Sbmlsim: SBML Simulation Made Easy. [object Object]. 10.5281/ZENODO.5531088
- Welsh, C., Xu, J., Smith, L., König, M., Choi, K., & Sauro, H. M. (2023). libRoadRunner 2.0: A High Performance SBML Simulation and Analysis Library. Bioinformatics, 39(1), btac770. 10.1093/bioinformatics/btac770
- Somogyi, E. T., Bouteiller, J.-M., Glazier, J. A., König, M., Medley, J. K., Swat, M. H., & Sauro, H. M. (2015). libRoadRunner: A High Performance SBML Simulation and Analysis Library. Bioinformatics, 31(20), 3315–3321. 10.1093/bioinformatics/btv363
- Smith, L. P., Hucka, M., Hoops, S., Finney, A., Ginkel, M., Myers, C. J., Moraru, I., & Liebermeister, W. (2015). SBML Level 3 Package: Hierarchical Model Composition, Version 1 Release 3. Journal of Integrative Bioinformatics, 12(2), 268. 10.2390/biecoll-jib-2015-268
- Bergmann, F. T., Adams, R., Moodie, S., Cooper, J., Glont, M., Golebiewski, M., Hucka, M., Laibe, C., Miller, A. K., Nickerson, D. P., Olivier, B. G., Rodriguez, N., Sauro, H. M., Scharm, M., Soiland-Reyes, S., Waltemath, D., Yvon, F., & Le Novère, N. (2014). COMBINE Archive and OMEX Format: One File to Share All Information to Reproduce a Modeling Project. BMC Bioinformatics, 15(1), 369. 10.1186/s12859-014-0369-z
- Bergmann, F. T., Rodriguez, N., & Le Novère, N. (2015). COMBINE Archive Specification Version 1. Journal of Integrative Bioinformatics, 12(2), 261. 10.2390/biecoll-jib-2015-261





